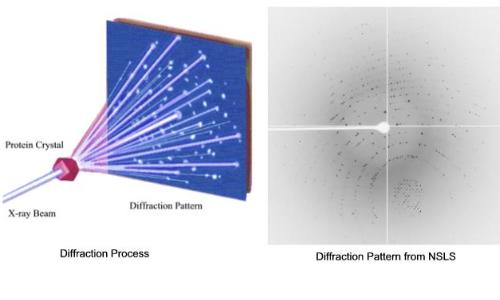
  3C, D image and diffraction pattern, respectively) appeared at about the same time (∼3 h) and was observed in vitrified specimens for several days after time zero. ), who, in an independent study, obtained similar crystal structures. The data of the latter two were recorded from a dry specimen, cooled to −180☌ to minimize electron beam radiation damage to the sensitive ChM crystals (see sample preparation). 2F shows the corresponding diffraction pattern. Figure 2E is an electron micrograph of such a crystal, and Fig. Both diffraction patterns are also identical to the diffraction pattern we obtained from a thin, standard ChM crystalline film, used as a reference, deposited on a carbon support film. The brightest ring closest to the center corresponds to 3.7 Å. 2D are produced by disordered crystalline ice together with vitreous ice. The d-spacing values of the diffraction patterns (two characteristic spots are marked on the image) agree with standard values to within 3% error, which can be attributed to measurement errors. 2B, D exhibit identical d-spacings and angles between reflections. This crystal had most likely begun to nucleate in the bile some time after time zero, because upon aspiration, samples are centrifuged and any existing crystals removed. Figure 2C shows a representative cryo-TEM image of a crystal seen in gallbladder bile number 2 ( Table 1), 3 days after removal from the patient, and Fig. The crystal habits were long, rectangular or classical, plate-like rhombus-shaped. Thus, a single representative image is shown. Several crystals were observed, and all exhibited identical diffraction patterns. The compositions and characteristics of these biles are shown in Table 1. Three native bile samples were studied by cryo-ED. Selected-area electron diffraction patterns were obtained from several crystals identified by imaging at low and medium magnifications of 3,400× and 40,000×, respectively. Images were recorded at nominal underfocus of 2–4 μm to enhance phase contrast. Singlecrystal electron diffraction pattern example software#A Gatan MultiScan791 cooled CCD camera operated with the Digital Micrograph 3.1 software package (Gatan), was used to acquire the images and diffraction patterns. Images and electron diffraction patterns were recorded in the low-dose mode to minimize electron beam radiation damage to the very radiation-sensitive samples. An Oxford CT3500 cooling-holder (Gatan Pleasanton, CA) was used to load the specimens into the TEM. Standard crystals (see above) that served as controls were prepared at room temperature and then cooled to liquid nitrogen temperature for sample stability. 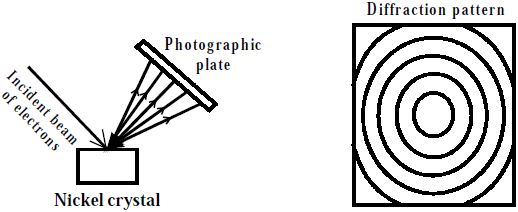
Thus, large crystals are always found flat on the grid. Structures larger than ∼300 nm in diameter are blotted off or may be distorted or positioned flat within the liquid film. We observed no evidence of anhydrous cholesterol crystallization in any of the biles studied. This crystal is exactly the monoclinic ChM phase described by Solomonov and coworkers ( Biophysical J., In press) in cholesterol monolayers compressed on the air–water interface. In solutions of model bile with low phospholipid-to-cholesterol ratio, electron diffraction provided direct proof of a novel transient polymorph that had an elongated habit and unit cell parameters differing from those of classic triclinic ChM. The growth and long-term stability of classic cholesterol monohydrate (ChM) crystals in native and model biles was determined. We combined cryogenic-temperature transmission electron microscopy with cryogenic-temperature electron diffraction to sequentially study crystal development and structure in nucleating model and native gallbladder biles. The exact structure of early-forming crystals is still controversial. Cholesterol crystals are the building blocks of cholesterol gallstones. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
